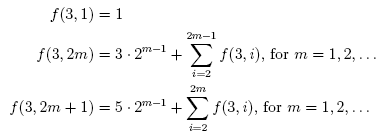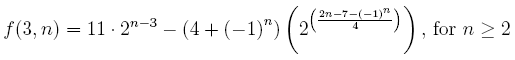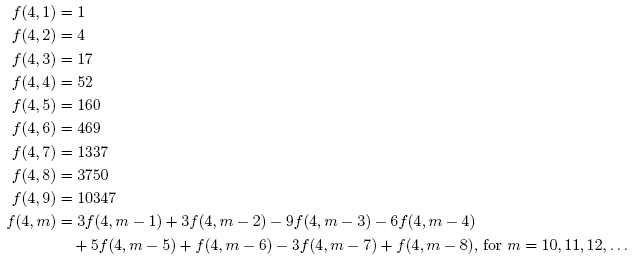There are a wide variety of different norms of matrices and operators that are useful in many different contexts. Some matrix norms, such as the Schatten norms and Ky Fan norms, are easy to compute thanks to the singular value decomposition. However, the computation of many other norms, such as the induced p-norms (when p ≠ 1, 2, ∞), is NP-hard. In this post, we will look at a general method for getting quite good estimates of almost any matrix norm.
The basic idea is that every norm can be written as a maximization of a convex function over a convex set (in particular, every norm can be written as a maximization over the unit ball of the dual norm). However, this maximization is often difficult to deal with or solve analytically, so instead it can help to write the norm as a maximization over two or more simpler sets, each of which can be solved individually. To illustrate how this works, let’s start with the induced matrix norms.
Induced matrix norms
The induced p → q norm of a matrix B is defined as follows:
\(\displaystyle\|B\|_{p\rightarrow q}:=\max\big\{\|B\mathbf{x}\|_q : \|\mathbf{x}\|_p = 1\big\},\)
where
\(\displaystyle\|\mathbf{x}\|_p := \left(\sum_{i}|x_i|^p\right)^{1/p}\)
is the vector p-norm. There are three special cases of these norms that are easy to compute:
- When p = q = 2, this is the usual operator norm of B (i.e., its largest singular value).
- When p = q = 1, this is the maximum absolute column sum: \(\|B\|_{1\rightarrow 1} = \max_j\sum_i|b_{ij}|\).
- When p = q = ∞, this is the maximum absolute row sum: \(\|B\|_{\infty\rightarrow \infty} = \max_i\sum_j|b_{ij}|\).
However, outside of these three special cases (and some other special cases, such as when B only has real entries that are non-negative [1]), this norm is much messier. In general, its computation is NP-hard [2], so how can we get a good idea of its value? Well, we rewrite the norm as the following double maximization:
\(\displaystyle\|B\|_{p\rightarrow q}=\max\big\{|\mathbf{y}^*B\mathbf{x}| : \|\mathbf{x}\|_p = 1, \|\mathbf{y}\|_{q^\prime} = 1\big\},\)
where \(q^\prime\) is the positive real number such that \(1/q + 1/q^\prime = 1\) (and we take \(q^\prime = 1\) if \(q = \infty\), and vice-versa). The idea is then to maximize over \(\mathbf{x}\) and \(\mathbf{y}\) one at a time, alternately.
- Start by setting \(j = 1\) and fixing a randomly-chosen vector \(\mathbf{x}_0\), scaled so that \(\|\mathbf{x}_0\|_{p} = 1\).
- Compute
\(\max\big\{|\mathbf{y}^*B\mathbf{x}_{j-1}| : \|\mathbf{y}\|_{q^\prime} = 1\big\},\)
keeping \(\mathbf{x}_{j-1}\) fixed, and let \(\mathbf{y}_j\) be the vector attaining this maximum. By Hölder’s inequality, we know that this maximum value is exactly equal to \(\|B\mathbf{x}_{j-1}\|_{q}\). Furthermore, the equality condition of Hölder’s inequality tells us that the vector \(\mathbf{y}_j\) attaining this maximum is the one with complex phases that are the same as those of \(B\mathbf{x}_{j-1}\), and whose magnitudes are such that \(|\mathbf{y}_j|^{q^\prime}\) is a multiple of \(|B\mathbf{x}_{j-1}|^q\) (here the notation \(|\cdot|^q\) means we take the absolute value and the q-th power of every entry of the vector).
- Compute
\(\max\big\{|\mathbf{y}_j^*B\mathbf{x}| : \|\mathbf{x}\|_{p} = 1\big\},\)
keeping \(\mathbf{y}_j\) fixed, and let \(\mathbf{x}_j\) be the vector attaining this maximum. By an argument almost identical to that of step 2, this maximum is equal to \(\|B^*\mathbf{y}_j\|_{p^\prime}\), where \(p^\prime\) is the positive real number such that \(1/p + 1/p^\prime = 1\). Furthermore, the vector \(\mathbf{x}_j\) attaining this maximum is the one with complex phases that are the same as those of \(B^*\mathbf{y}_j\), and whose magnitudes are such that \(|\mathbf{x}_j|^p\) is a multiple of \(|B^*\mathbf{y}_j|^{p^\prime}\).
- Increment \(j\) by 1 and return to step 2. Repeat until negligible gains are made after each iteration.
This algorithm is extremely quick to run, since Hölder’s inequality tells us exactly how to solve each of the two maximizations separately, so we’re left only performing simple vector calculations at each step. The downside of this algorithm is that, even though it will always converge to some local maximum, it might converge to a value that is smaller than the true induced p → q norm. However, in practice this algorithm is fast enough that it can be run several thousand times with different (randomly-chosen) starting vectors \(\mathbf{x}_0\) to get an extremely good idea of the value of \(\|B\|_{p\rightarrow q}\).
It is worth noting that this algorithm is essentially the same as the one presented in [3], and reduces to the power method for finding the largest singular value when p = q = 2. This algorithm has been implemented in the QETLAB package for MATLAB as the InducedMatrixNorm function.
Induced Schatten superoperator norms
There is a natural family of induced norms on superoperators (i.e., linear maps \(\Phi : M_n \rightarrow M_n\)) as well. First, for a matrix \(X \in M_n\), we define its Schatten p-norm to be the p-norm of its vector of singular values:
\(\|X\|_p := \left(\sum_{i=1}^n \sigma_i(X)^p\right)^{1/p}.\)
Three special cases of the Schatten p-norms include:
- p = 1, which is often called the “trace norm” or “nuclear norm”,
- p = 2, which is often called the “Frobenius norm” or “Hilbert–Schmidt norm”, and
- p = ∞, which is the usual operator norm.
The Schatten norms themselves are easy to compute (since singular values are easy to compute), but their induced counter-parts are not.
Given a superoperator \(\Phi : M_n \rightarrow M_n\), its induced Schatten p → q norm is defined as follows:
\(\|\Phi\|_{p\rightarrow q} := \max\big\{ \|\Phi(X)\|_q : \|X\|_p = 1 \big\}.\)
These induced Schatten norms were studied in some depth in [4], and crop up fairly frequently in quantum information theory (especially when p = q = 1) and operator theory (especially when p = q = ∞). The fact that they are NP-hard to compute in general is not surprising, since they reduce to the induced matrix norms (discussed earlier) in the case when \(\Phi\) only acts on the diagonal entries of \(X\) and just zeros out the off-diagonal entries. However, it seems likely that this norm’s computation is also difficult even in the special cases p = q = 1 and p = q = ∞ (however, it is straightforward to compute when p = q = 2).
Nevertheless, we can obtain good estimates of this norm’s value numerically using essentially the same method as discussed in the previous section. We start by rewriting the norm as a double maximization, where each maximization individually is easy to deal with:
\(\|\Phi\|_{p\rightarrow q} = \max\big\{ |\mathrm{Tr}(Y^*\Phi(X))| : \|X\|_p = 1, \|Y\|_{q^\prime} = 1\big\},\)
where \(q^\prime\) is again the positive real number (or infinity) satisfying \(1/q + 1/q^\prime = 1\). We now maximize over \(X\) and \(Y\), one at a time, alternately, just as before:
- Start by setting \(j = 1\) and fixing a randomly-chosen matrix \(X_0\), scaled so that \(\|X_0\|_p = 1\).
- Compute
\(\max\big\{|\mathrm{Tr}(Y^*\Phi(X_{j-1})| : \|Y\|_{q^\prime} = 1\big\},\)
keeping \(X_{j-1}\) fixed, and let \(Y_j\) be the matrix attaining this maximum. By the Hölder inequality for Schatten norms, we know that this maximum value is exactly equal to \(\|\Phi(X_{j-1})\|_{q}\). Furthermore, the matrix \(Y_j\) attaining this maximum is the one with the same left and right singular vectors as \(\Phi(X_{j-1})\), and whose singular values are such that there is a constant \(c\) so that \(\sigma_i(Y_j)^{q^\prime} = c\sigma_i(\Phi(X_{j-1}))^q\) for all \(i\) (i.e., the vector of singular values of \(Y_j\), raised to the \(q^\prime\) power, is a multiple of the vector of singular values of \(\Phi(X_{j-1})\), raised to the \(q\) power).
- Compute
\(\max\big\{|\mathrm{Tr}(Y_j^*\Phi(X)| : \|X\|_{p} = 1\big\},\)
keeping \(Y_j\) fixed, and let \(X_j\) be the matrix attaining this maximum. By essentially the same argument as in step 2, we know that this maximum value is exactly equal to \(\|\Phi^*(Y_j)\|_{p^\prime}\), where \(\Phi^*\) is the map that is dual to \(\Phi\) in the Hilbert–Schmidt inner product. Furthermore, the matrix \(X_j\) attaining this maximum is the one with the same left and right singular vectors as \(\Phi^*(Y_j)\), and whose singular values are such that there is a constant \(c\) so that \(\sigma_i(X_j)^{p} = c\sigma_i(\Phi^*(Y_j))^{p^\prime}\) for all \(i\).
- Increment \(j\) by 1 and return to step 2. Repeat until negligible gains are made after each iteration.
The above algorithm is almost identical to the algorithm presented for induced matrix norms, but with absolute values and complex phases of the vectors \(\mathbf{x}\) and \(\mathbf{y}\) replaced by the singular values and singular vectors of the matrices \(X\) and \(Y\), respectively. The entire algorithm is still extremely quick to run, since each step just involves computing one singular value decomposition.
The downside of this algorithm, as with the induced matrix norm algorithm, is that we have no guarantee that this method will actually converge to the induced Schatten p → q norm; only that it will converge to some lower bound of it. However, the algorithm works pretty well in practice, and is fast enough that we can simply run it a few thousand times to get a very good idea of what the norm actually is. If you’re interested in making use of this algorithm, it has been implemented in QETLAB as the InducedSchattenNorm function.
Entanglement Norms
The central idea used for the previous two families of norms can also be used to get lower bounds on the following norm on \(M_m \otimes M_n\) that comes up from time to time when dealing with quantum entanglement:
\(\|X\|_{S(1)} := \max\Big\{\big|(\mathbf{v}\otimes\mathbf{w})^*X(\mathbf{x} \otimes \mathbf{y})\big| : \|\mathbf{v}\| = \|\mathbf{w}\| = \|\mathbf{x}\| = \|\mathbf{y}\| = 1\Big\}.\)
(As a side note: this norm, and some other ones like it, were the central focus on my thesis.) This norm is already written for us as a double maximization, so the idea presented in the previous two sections is somewhat clearer from the start: we fix randomly-generated vectors \(\mathbf{v}\) and \(\mathbf{x}\) and then maximize over all vectors \(\mathbf{w}\) and \(\mathbf{y}\), which can be done simply by computing the left and right singular vectors associated with the maximum singular value of the operator
\((\mathbf{v} \otimes I)^*X(\mathbf{x} \otimes I) \in M_n.\)
We then fix \(\mathbf{w}\) and \(\mathbf{y}\) as those singular vectors and then maximize over all vectors \(\mathbf{v}\) and \(\mathbf{x}\) (which is again a singular value problem), and we iterate back and forth until we converge to some value.
As with the previously-discussed norms, this algorithm always converges, and it converges to a lower bound of \(\|X\|_{S(1)}\), but perhaps not its exact value. If you want to take this algorithm out for a spin, it has been implemented in QETLAB as the sk_iterate function.
It’s also worth mentioning that this algorithm generalizes straightforwardly in several different directions. For example, it can be used to find lower bounds on the norms \(\|\cdot\|_{S(k)}\) where we maximize on the left and right by pure states with Schmidt rank not larger than k rather than separable pure states, and it can be used to find lower bounds on the geometric measure of entanglement [5].
References:
- D. Steinberg. Computation of matrix norms with applications to robust optimization. Research thesis. Technion – Israel University of Technology, 2005.
- J. M. Hendrickx and A. Olshevsky. Matrix p-norms are NP-hard to approximate if p ≠ 1,2,∞. 2009. E-print: arXiv:0908.1397
- D. W. Boyd. The power method for ℓp norms. Linear Algebra and Its Applications, 9:95–101, 1974.
- J. Watrous. Notes on super-operator norms induced by Schatten norms. Quantum Information & Computation, 5(1):58–68, 2005. E-print: arXiv:quant-ph/0411077
- T.-C. Wei and P. M. Goldbart. Geometric measure of entanglement and applications to bipartite and multipartite quantum states. Physical Review A, 68:042307, 2003. E-print: arXiv:quant-ph/0212030









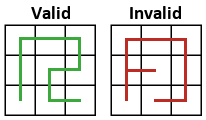 Before proceeding, let’s simplify things slightly by making the restriction that the maximal self-avoiding walks must start in the bottom-l
Before proceeding, let’s simplify things slightly by making the restriction that the maximal self-avoiding walks must start in the bottom-l